Suppose that you've estimated an econometric model and you want to test the residuals for serial independence, or perhaps for homoskedasticity. The trouble is that for the model and estimator that you've used, your favourite computer package doesn't provide such tests. Is there a quick way of "tricking" the package into giving you the information that you want?
Yes, there is. I'll show you how, by looking at a situation that arises when using EViews. Similar examples occur with pretty much any econometrics package that you happen to be using, so the idea here has wide applicability.
If you use EViews to estimate a "Seemingly Unrelated Regression Equations" (SURE) model, then you'll find that you have very few options when it comes to conducting diagnostic tests on the residuals of the equations in the system. For example, no tests for the homoskedasticity of the errors are immediately available. When you run into this sort of problem, here's one way that you can "fool" the package into giving you access to some of the tests you want, with minimal effort.
The trick is to be aware of the following result:
To see this, consider the following "artificial" regression:
yi = γ + εi ; i = 1, 2. ...., n.
The OLS estimator of gamma is: c = Σ (yi 1) / Σ (1) = Σ (yi) / n.
Then, the residuals are ei = yi - Σ (yi) / n. In particular, if the "y" data had a mean of zero to begin with, then ei = yi ; for all i.
So, if we have a residuals series from some estimated model, and we then use this series as the y variable in the above artificial regression, any tests that we then perform on the latter regression's residuals will actually be tests on the y series, or on the residuals from the original model that we estimated. In this way we can get access to any tests available through the OLS command and apply them to the residuals from the other model.
Of course, if we did this, we'd need to make sure that the tests in question were valid when applied to these other residuals.
I have an example using monthly data for the market shares of various web browsers between October 2004 and February 2007. The data are available on the Data page that accompanies this blog. The example involves 6-equation SURE model for these market shares. If you look at the data files, you'll see the definitions of the variables. The EViews workfile that I've used or the following analysis is in the Code page for this blog.
Here is the specification of my model:
and the rest of the output is:
The residuals for each equation sum to zero, as can been in the following table:
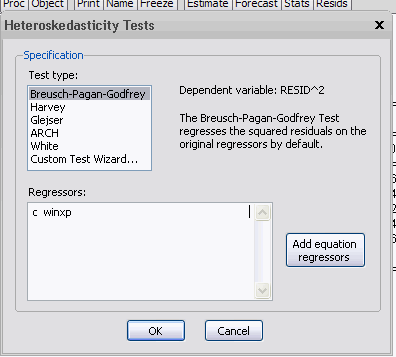
The residuals from the regression are actually just RESID01 - the residuals series for the first equation in the SURE system. I can now select "VIEW / RESIDUAL DIAGNOSTICS / HETEROSKEDASTICITY TESTS", and I get:
Not all of these tests will be appropriate. For example, I can't use White's test because it will simply use the "regressors" from this artificial regression (namely a column of ones), and their squared values. However, the B-P-G, Harvey, and Glesjer tests can all be used. If, for example, I use the B-P-G test, and specify that the heteroskedasticity may be a function of WINXP, I get the following results:
The p-value associated with the nR2 version of the test suggests that we should reject the null hypothesis of homoskedasticity at the 3.7% significance level (or higher).
If we apply the same test to the residuals of the fifth equation, postulating that any heteroskedasticity may be associated with the WIN2000 variable, we get:
The trick is to be aware of the following result:
If we fit a regression equation in which the only "regressor" is a column of ones, then the residuals from this curious regression will just be the data for the dependent variable, adjusted so that they are in deviations about the sample mean.
To see this, consider the following "artificial" regression:
yi = γ + εi ; i = 1, 2. ...., n.
The OLS estimator of gamma is: c = Σ (yi 1) / Σ (1) = Σ (yi) / n.
Then, the residuals are ei = yi - Σ (yi) / n. In particular, if the "y" data had a mean of zero to begin with, then ei = yi ; for all i.
So, if we have a residuals series from some estimated model, and we then use this series as the y variable in the above artificial regression, any tests that we then perform on the latter regression's residuals will actually be tests on the y series, or on the residuals from the original model that we estimated. In this way we can get access to any tests available through the OLS command and apply them to the residuals from the other model.
Of course, if we did this, we'd need to make sure that the tests in question were valid when applied to these other residuals.
I have an example using monthly data for the market shares of various web browsers between October 2004 and February 2007. The data are available on the Data page that accompanies this blog. The example involves 6-equation SURE model for these market shares. If you look at the data files, you'll see the definitions of the variables. The EViews workfile that I've used or the following analysis is in the Code page for this blog.
Here is the specification of my model:
The estimated coefficients are as follows:
Now, to illustrate the idea outlined above, I've tested the residuals of each equation for homoskedasticity. Let's consider the first equation. First, I estimate the artificial regression by OLS:
If we apply the same test to the residuals of the fifth equation, postulating that any heteroskedasticity may be associated with the WIN2000 variable, we get:
The small p-value again suggests that we should reject the null hypothesis of homoskedasticity for the errors of this equation of the original SURE model.
Two further comments are in order:
- It's worth re-emphasizing that any tests that we perform in this way must be applicable to the residuals being analyzed. Here, the residuals were for the equations of a SURE model. The homoskedasticity tests being applied are asymptotically valid tests, so they will be relevant in this case. Our sample size is small, however, so the result should be treated with caution, just as they should be if we had applied them to an original OLS regression with this sample size.
- Having detected the presence of heteroskedasticity, we could now transform the data for the variables in the equations in question, and then re-estimate the SURE model. In other words, we could modify the SURE estimation in a Weighted Least Squares manner.
© 2012, David E. Giles







Hey, Prof, won't tests implemented in this fashion be optimistic in terms of the degrees of freedom they associated with the estimated residuals? If you have n residuals from your original fitted model which estimated, say, k parameters, then we should be using n-k as our residual degrees of freedom but presumably this method assumes the residuals from the second model (which are equivalent to those from the first) have n-1 degrees of freedom (or possibly n if a no intercept / no regressor model is fit). So won't some tests be off?
ReplyDeleteBen - thanks for the comment. Keep in mind that these LM tests have only asymptotic validity, so there is no distinction between n and (n-k). Bottom line: no real problem.
DeleteHi Prof. Thanks for the insights. However, suppose you need to do a multivariate test of heteroskedasticity for the whole SUR system rather than equation-by-equation. How do you do it in Eviews? Kindly refer to the following paper:
ReplyDeleteSystem Misspecification Testing and Structural Change in the Demand for Meats
Anya McGuirk, Paul Driscoll, Jeffrey Alwang, and Huilin Huang (1995).
There's need to take into account cross-equation correlation when conducting multivariate diagnostic tests.